Main menu
You are here
Tutorial 6. Data categories
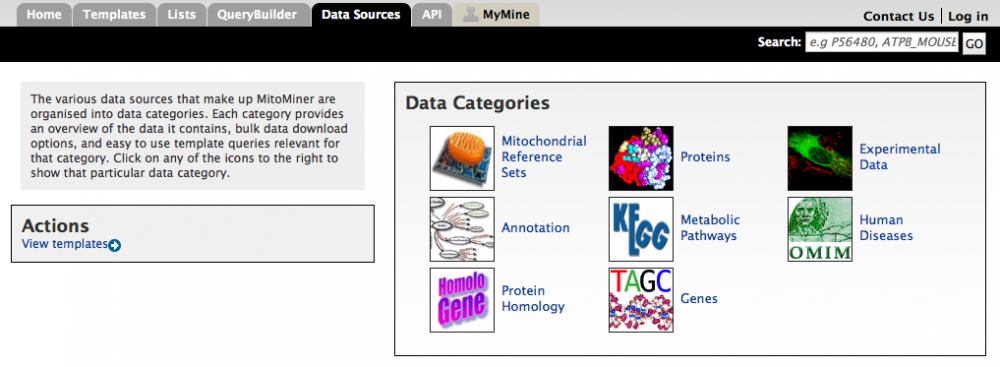
The various data sources that make up MitoMiner are organised into data categories. These categories can be accessed from the 'Data Sources' tab on the main menu bar.
Each data category page consists of four different sections:
1. The first section provides a brief overview and description of the data category, and information such as when it was last updated and the type of data included.
2. The second section is configured to provide bulk download options for the data source. MitoMiner allows this data to be exported as a comma or tab delimited values, which is compatible with most spreadsheet programs including MS Excel.
3. The third section contains a selection of template queries (described in tutorial 3) that are relevant to that data source. These queries are envisaged to be the most common queries required by a typical user. These template queries can be run "as is" or can be further modified to meet particular requirements by using the query builder.
4. The fourth section is configured to provide pertinent starting points to create new queries from scratch using the query builder.
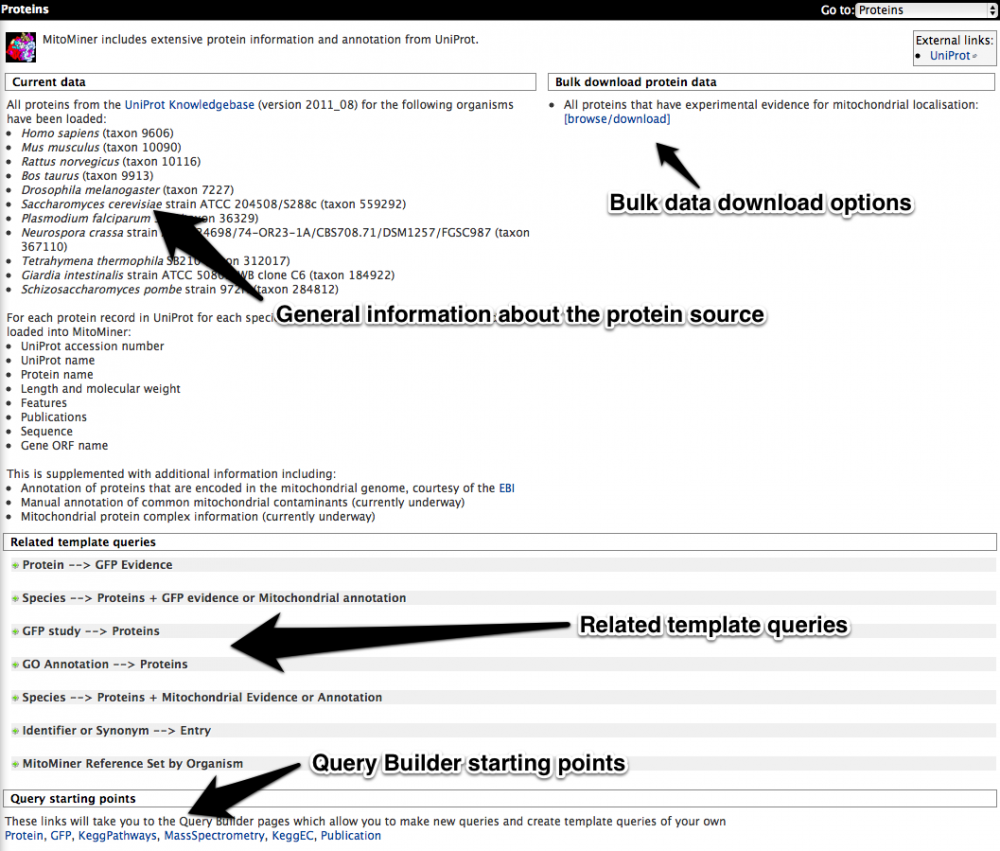
As an example the Protein data category is shown below. The first section provides details such as the version of UniProt used and the various different species loaded. The second section allows the bulk download of "all proteins that have experimental evidence of mitochondrial localisation". The third section contains relevant template queries such as "Proteins --> Mitochondrial Evidence or Annotation". The fourth section contains aspect relevant starting points for the query builder such as protein or GFP.